Go Big To Go Home
The first step of data science when working with new a dataset is to understand the high-level facts and relationships within the data. This is often done by exploring the data interactively by using something like Python, R, or Matlab.
Recently I've been exploring a new dataset. It's a pretty big dataset: a few hundred gigabytes of data in compressed Parquet format. A rule of thumb is that reading data off disk into memory takes ten or twenty times the memory than the storage the data uses on disk. For this dataset, that could equal more than ten terabytes of memory, which in 2025 is still a pretty ridiculous amount of memory on a single machine. It is for this reason that working with data this size requires tools that allow you to work with the data without loading all of it into memory at once.
One of these tools is Polars. Quoting the Polars home page: "Want to process large data sets that are bigger than your memory? Our streaming API allows you to process your results efficiently, eliminating the need to keep all data in memory." It's still a bit rough around the edges with some unfinished and missing features, but overall it's a powerful and capable tool for data analysis. Lately I've been using Polars more and more, taking advantage of this "streaming" ability.
On the other hand, sometimes it's easiest to do things directly and skip all the low-memory "streaming" tricks. If I can get an answer more quickly by simply using lots of memory, especially if it's something I'm doing only once and not putting into a repeated process, then this can be the right choice. Polars can do "streaming" analysis, but at a certain point it has to coalesce things into an answer, and sometimes that answer can use a significant amount of memory.
There are many negative aspects of cloud computing that I won't get into here. However, there are some good things, and one of them is that you can scale up and down resources as needed. All cloud providers, like Amazon Web Services and Google Cloud Platform offer many services and in particular Virtual Machines. When running a virtual machine you can choose the hardware specifications in terms of CPU kind and core count, amount of RAM, and other features like network speeds, GPUs, or SSDs. A virtual machine can be booted on one hardware configuration, shut down, and the rebooted on a different configuration as needed. It's as if you took the hard drive out of your laptop and put it in a big workstation. All your data and settings are still there, but you've upgraded the hardware. This is something I take advantage of frequently!
I was attempting to do a certain analysis of the new dataset on an EC2 virtual machine and I kept running out of memory. Instead of switching to some low-memory tricks, I decided to see if I could save some time by simply rebooting my virtual machine on one of the larger instances AWS offers: a r7i.48xlarge. This has 192 CPU cores and 1,536 GB of RAM. It costs $12.70 per hour. I get paid more than $12.70 per hour, so if booting up a huge machine like this saves me even a little time, it's worth it.
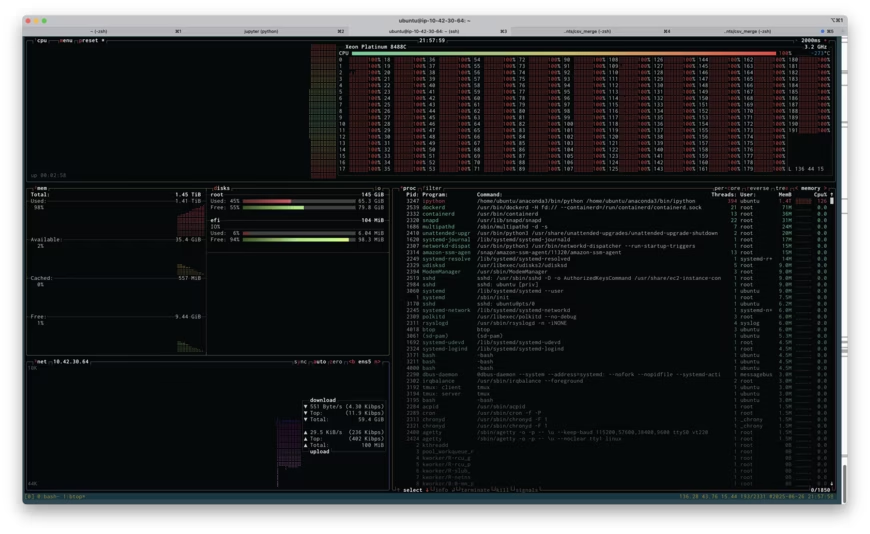
Above is a screenshot of btop running on the r7i.48xlarge instance while I attempted to run my analysis. If you click on the image, you'll see the full size screenshot. You'll see that I'm about to run out of memory: 1.41TB used of 1.45TB. You'll also see that I'm using all 192 cores at 100% load (the cores are labeled 0-191). Unfortunately, throwing all this memory at the problem didn't work, I ran out of memory, and I had to resort to being more clever. Being clever took more time, of course, but if the high-RAM instance had worked, it would have paid off.
Playing with various server configuration tools (like this one) shows that the r7i.48xlarge would cost at least $60,000. This is not something that I need very often, and purchasing something this large would be ridiculous. However, renting it for half an hour, if it saves me a few hours, is definitely worth it. Also, it's kind of fun to say "yeah, I used 1.5TB of memory and 192 cores and it wasn't enough."
more ...Polars scan_csv and sink_parquet
The documentation for polars is not the best,
and figuring out how to do this below took me over an hour.
Here's how to read in a headerless csv file into a
LazyFrame using
scan_csv
and write it to a parquet file using
sink_parquet.
The key is to use with_column_names and
schema_overrides.
Despite what the documentation says, using schema doesn't work as you
might imagine and sink_parquet returns with a cryptic error about
the dataframe missing column a.
This is just a simplified version of what I actually am trying to do, but that's the best way to drill down to the issue. Maybe the search engines will find this and save someone else an hour of frustration.
import numpy as np
import polars as pl
df = pl.DataFrame(
{"a": [str(i) for i in np.arange(10)], "b": np.random.random(10)},
)
df.write_csv("/tmp/stuff.csv", include_header=False)
lf = pl.scan_csv(
"/tmp/stuff.csv",
has_header=False,
schema_overrides={
"a": pl.String,
"b": pl.Float64,
},
with_column_names=lambda _: ["a", "b"],
)
lf.sink_parquet("/tmp/stuff.parquet")